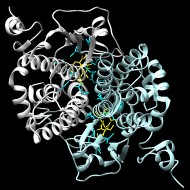
The Main subgroup of cytosolic GSTs (cytGSTs) is a large and diverse class of cytGSTs that encompasses the phi, tau, theta, HSP26, zeta, omega, and beta cytGSTs. While most of these GSTs are bacterial, this group includes large numbers of fungi, animal, and plant GSTs as well. Some of these members use a catalytic serine to activate glutathione, and others use a cysteine. The cysteine of omega class GSTs is in the same position relative to the fold as the serines of phi, tau, theta, HSP26, and zeta classes, while the beta GSTs have a cysteine two positions later. A minority of Main subgroup cytGSTs (including E. coli YfcG) apparently use a threonine to activate glutathione.
Armstrong RN
Structure, catalytic mechanism, and evolution of the glutathione transferases
▸ Abstract
None
Chem Res Toxicol
1997;10(1):2-18
| PubMed ID:
9074797
Jemth P, Mannervik B
Active site serine promotes stabilization of the reactive glutathione thiolate in rat glutathione transferase T2-2. Evidence against proposed sulfatase activity of the corresponding human enzyme
▸ Abstract
Ser(11) in rat glutathione transferase T2-2 is important for stabilization of the reactive enzyme-bound glutathione thiolate in the reaction with 1-menaphthyl sulfate. The S11A mutation increased the pK(a) value for the pH dependence of the rate constant for pre-steady-state product formation, from 5.7 to 7.9. This pH dependence is proposed to reflect titration of enzyme-bound glutathione thiol. Further, the mutation lowered the k(cat) value but not because of the impaired stabilization of the glutathione thiolate. In fact, several steps on the reaction pathway were affected by the S11A mutation, and the cause of the decreased k(cat) for the mutant was found to be a slower product release. The data presented here contradict the hypothesis that glutathione transferase T2-2 could act as a sulfatase that is not dependent on Ser(11) for the catalytic activity, as proposed for the corresponding human enzyme (Tan, K.-L., Chelvanayagam, G., Parker, M. W., and Board, P. G. (1996) Biochem. J. 319, 315-321; Rossjohn, J., McKinstry, W. J., Oakley, A. J., Verger, D., Flanagan, J., Chelvanayagam, G., Tan, K.-L., Board, P. G., and Parker, M. W. (1998) Structure 6, 309-322). On the contrary, Ser(11) governs both chemical and physical steps of the catalyzed reaction.
J Biol Chem
2000;275(12):8618-8624
| PubMed ID:
10722701
Copley SD
Evolution of a metabolic pathway for degradation of a toxic xenobiotic: the patchwork approach
▸ Abstract
The pathway for degradation of the xenobiotic pesticide pentachlorophenol in Sphingomonas chlorophenolica probably evolved in the past few decades by the recruitment of enzymes from two other catabolic pathways. The first and third enzymes in the pathway, pentachlorophenol hydroxylase and 2,6-dichlorohydroquinone dioxygenase, may have originated from enzymes in a pathway for degradation of a naturally occurring chlorinated phenol. The second enzyme, a reductive dehalogenase, may have evolved from a maleylacetoacetate isomerase normally involved in degradation of tyrosine. This apparently recently assembled pathway does not function very well: pentachlorophenol hydroxylase is quite slow, and tetrachlorohydroquinone dehalogenase is subject to severe substrate inhibition.
Trends Biochem Sci
2000;25(6):261-265
| PubMed ID:
10838562
Sheehan D, Meade G, Foley VM, Dowd CA
Structure, function and evolution of glutathione transferases: implications for classification of non-mammalian members of an ancient enzyme superfamily
▸ Abstract
The glutathione transferases (GSTs; also known as glutathione S-transferases) are major phase II detoxification enzymes found mainly in the cytosol. In addition to their role in catalysing the conjugation of electrophilic substrates to glutathione (GSH), these enzymes also carry out a range of other functions. They have peroxidase and isomerase activities, they can inhibit the Jun N-terminal kinase (thus protecting cells against H(2)O(2)-induced cell death), and they are able to bind non-catalytically a wide range of endogenous and exogenous ligands. Cytosolic GSTs of mammals have been particularly well characterized, and were originally classified into Alpha, Mu, Pi and Theta classes on the basis of a combination of criteria such as substrate/inhibitor specificity, primary and tertiary structure similarities and immunological identity. Non-mammalian GSTs have been much less well characterized, but have provided a disproportionately large number of three-dimensional structures, thus extending our structure-function knowledge of the superfamily as a whole. Moreover, several novel classes identified in non-mammalian species have been subsequently identified in mammals, sometimes carrying out functions not previously associated with GSTs. These studies have revealed that the GSTs comprise a widespread and highly versatile superfamily which show similarities to non-GST stress-related proteins. Independent classification systems have arisen for groups of organisms such as plants and insects. This review surveys the classification of GSTs in non-mammalian sources, such as bacteria, fungi, plants, insects and helminths, and attempts to relate them to the more mainstream classification system for mammalian enzymes. The implications of this classification with regard to the evolution of GSTs are discussed.
Biochem J
2001;360(None):1-16
| PubMed ID:
11695986
Wadington MC, Ladner JE, Stourman NV, Harp JM, Armstrong RN
Analysis of the structure and function of YfcG from Escherichia coli reveals an efficient and unique disulfide bond reductase
▸ Abstract
YfcG is one of eight glutathione (GSH) transferase homologues encoded in the Escherichia coli genome. The protein exhibits low or no GSH transferase activity toward a panel of electrophilic substrates. In contrast, it has a very robust disulfide-bond reductase activity toward 2-hydroxyethyldisulfide on par with mammalian and bacterial glutaredoxins. The structure of YfcG at 2.3 A-resolution from crystals grown in the presence of GSH reveals a molecule of glutathione disulfide in the active site. The crystallographic results and the lack of functional cysteine residues in the active site of YfcG suggests that the reductase activity is unique in that no sulfhydryl groups in the YfcG protein are covalently involved in the redox chemistry.
Biochemistry
2009;48(28):6559-6561
| PubMed ID:
19537707