| Total |
100%  |
<100%  |
Family unknown  |
||||
| Functional domains | 401 | ||||||
| UniProtKB | 522 | 0 | 0 | 522 | |||
| GI | 1561 | 0 | 0 | 1561 | |||
| Structures | 15 | ||||||
| Reactions | 0 | ||||||
| Functional domains of this subgroup were last updated on June 10, 2017 | |||||||
| New functional domains were last added to this subgroup on June 22, 2012 | |||||||
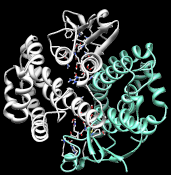
Cytosolic GSTs that include proteins with "Phi" Swiss-Prot family annotations. Proteins in this subgroup are known to catalyze the following reaction types: epoxide ring opening, nucleophilic aromatic substitution, and peroxidase reactions. For example, SFLD protein 58579 (UniProt accn P46422) has experimentally-confirmed nucleophilic aromatic substitution activity with CDNB and epoxide ring opening activity with 1,2-epoxy-3-(p-nitrophenoxy)propane.
No References.
No notes.
Static File Downloads
| File Name | Description | Parameters | Stats |
|---|---|---|---|
| sfld_alignment_sg1154.msa | Annotated Sequence Alignment, Stockholm format | 58 sequences size: 16K |
Total number of functional domains in this group.
Number of Functional Domains that have been manually or automatically been assigned to a family.
Number of Functional Domains that have not been assigned to a family.
Number of structures available from the PDB for members of this group.
Number of Functional Domains with 100% of Conserved Residues
Number of Functional Domains with less than 100% Conserved Residues
| Subgroup ▸ Legend | T  |
S  |
||
|---|---|---|---|---|
| ┗ 1: Main.5: Phi-like | 401 | 15 |
Depth of the multi-level Subgroup hierarchy.