Top Level Name
⌊ Superfamily (core) Haloacid Dehalogenase
⌊ Subgroup C1.5: HAD, Beta-PGM, Phosphatase Like
⌊ C1.5.5: Heptose Bisphosphate Phosphatase Like
⌊ Family histidinol-phosphatase
| Total |
100%  |
<100%  |
|||
| Functional domains | 3 | 0 | 3 | ||
| UniProtKB | 37 | 0 | 37 | ||
| GI | 90 | 0 | 90 | ||
| Structures | 0 | ||||
| Reactions | 1 | ||||
| Functional domains of this family were last updated on Nov. 22, 2017 | |||||
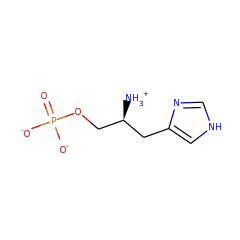
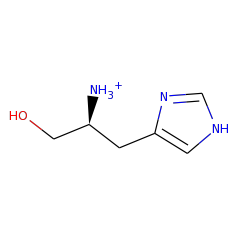
Enzymes in the histidinol-phosphatase family contain two structural domains, each with a seperate active site. Only the n-terminal domain is part of the haloacid dehalogenase superfamily. This domain catalyzes the dephosphorylation of L-histidinol-phosphate to L-histidinol. The c-terminal domain of the characterized E. coli enzyme has imidazoleglycerol-phosphate dehydratase activity, but is not part of the HAD superfamily.
Chiariotti, L., et al.
Gene structure in the histidine operon of Escherichia coli. Identification and nucleotide sequence of the hisB gene
▸ Abstract
Mol Gen Genet 1986;202(1):42-47 | PubMed ID: 3007936
No notes.
Static File Downloads
| File Name | Description | Parameters | Stats |
|---|---|---|---|
| sfld_alignment_fam28.msa | Annotated Sequence Alignment, Stockholm format | 2 sequences size: 1.4K |