Top Level Name
⌊ Superfamily (core) Haloacid Dehalogenase
⌊ Subgroup C1.5: HAD, Beta-PGM, Phosphatase Like
⌊ C1.5.6: HAD, Beta-PGM, Phosphatase Like
⌊ FunctionalDomain C1.5.6: HAD, Beta-PGM, Phosphatase Like (ID 126304)
No Notes.
| Superfamily Assignment Evidence Code(s) | ISS |
| This entry was last updated on | Nov. 22, 2017 |
References to Other Databases
Genbank
| Species | GI | Accession | Proteome |
|---|---|---|---|
| Homo sapiens Taxon ID: 9606 | 23308749 | NP_689880.1 (RefSeq) | PRP URP |
| Gorilla gorilla gorilla Taxon ID: 9595 | 426391228 | XP_004061981.1 (RefSeq) | URP |
| synthetic construct Taxon ID: 32630 | 649114641 | AIC53137.1 (Genbank) | |
| synthetic construct Taxon ID: 32630 | 312150668 | ADQ31846.1 (Genbank) | |
| synthetic construct Taxon ID: 32630 | 190691161 | ACE87355.1 (Genbank) | |
| synthetic construct Taxon ID: 32630 | 190689793 | ACE86671.1 (Genbank) | |
| Homo sapiens Taxon ID: 9606 | 119630486 | EAX10081.1 (Genbank) | PRP URP |
| n/a | 91130759 | ABE14507.1 (Genbank) | |
| n/a | 75921233 | ABA32504.1 (Genbank) | |
| Homo sapiens Taxon ID: 9606 | 18490374 | AAH22552.1 (Genbank) | PRP URP |
| Homo sapiens Taxon ID: 9606 | 193786241 | PRP URP | |
| Homo sapiens Taxon ID: 9606 | 30315932 | PRP URP | |
| obsolete GI = 56203139 | |||
| Show All | |||
Uniprot
| Protein Name | Accession | EC Number
 |
Identifier |
|---|---|---|---|
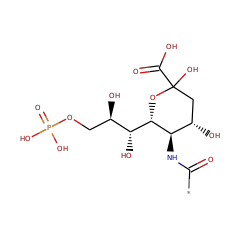
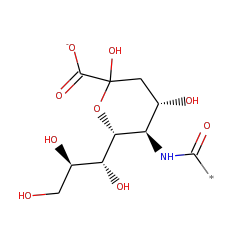
| N-acylneuraminate-9-phosphatase | Q8TBE9 | 3.1.3.29 | NANP_HUMAN (Swiss-Prot) |
Length of Enzyme (full-length): 248 | Length of Functional Domain: 238
MGLSRVRAVFFDLDNTLIDTAGASRRGMLEVIKLLQSKYHYKEEAEIICDKVQVKLSKEC
FHPYNTCITDLRTSHWEEAIQETKGGAANRKLAEECYFLWKSTRLQHMTLAEDVKAMLTE
LRKEVRLLLLTNGDRQTQREKIEACACQSYFDAVVVGGEQREEKPAPSIFYYCCNLLGVQ
PGDCVMVGDTLETDIQGGLNAGLKATVWINKNGIVPLKSSPVPHYMVSSVLELPALLQSI
DCKVSMST
Conserved catalytic residues (as determined by automated alignment to family, subgroup, or superfamily HMMs) are shown with teal highlighting . Conserved catalytic residues which do not matched the Conserved Alignment Residue are shown with maroon highlighting . Information regarding their function can be found in the Conserved Residues section below.