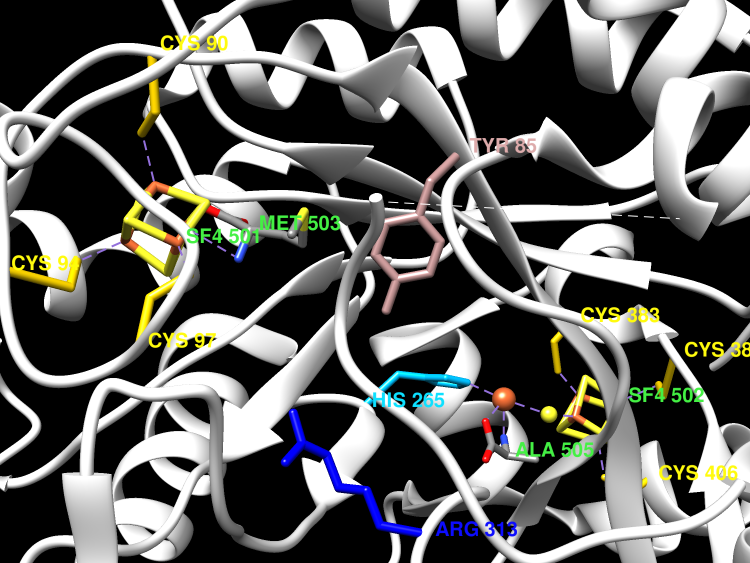
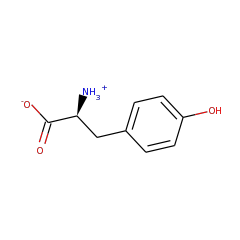
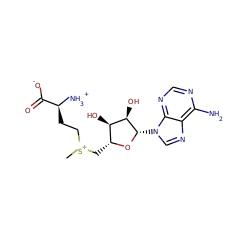
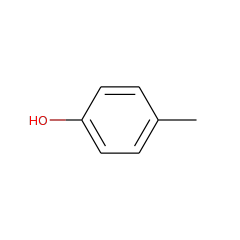
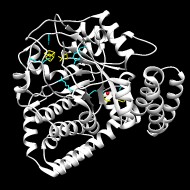
This is one of the three enzymes required for the formation of the [FeFe]-hydrogenase metallocofactor. Recent work suggests that HydG is a multifunctional enzyme that produces tyrosine-derived dehydroglycine (DHG), CN(-), and CO at three well-differentiated catalytic sites.
Given the sequence homology between HydG and ThiH, it is proposed that HydG is specifically involved in the formation of the dithiolate ligand portion of the metallocofactor (with HydE being responsible for the carbon monoxide and cyanate ligands) with two DHG molecules as precursors along with an [FeS] cluster of HydG functioning both as electron shuttle and source of the sulfur atoms.
Recent work by Kuchenreuther et al. suggests that the radical SAM reaction, initiated at an N-terminal 4Fe-4S cluster, generates a tyrosine radical bound to a C-terminal 4Fe-4S cluster. Heterolytic cleavage of this tyrosine radical at the Calpha-Cbeta bond forms a transient 4-oxidobenzyl radical and a dehydroglycine bound to the C-terminal 4Fe-4S cluster. Dinis et al. have shown that this C-terminal bound iron-sulfur cluster is initially [5Fe-5S] and that the fifth labile iron is the site of the Fe(CO)x(CN)y synthon formation.
Pilet E, Nicolet Y, Mathevon C, Douki T, Fontecilla-Camps JC, Fontecave M
The role of the maturase HydG in [FeFe]-hydrogenase active site synthesis and assembly.
▸ Abstract
[FeFe]-hydrogenases catalyze the protons/hydrogen interconversion through a unique di-iron active site consisting of three CO and two CN ligands, and a non-protein SCH(2)XCH(2)S (X=N or O) dithiolate bridge. Site assembly requires two "Radical-S-adenosylmethionine (SAM or AdoMet)" iron-sulfur enzymes, HydE and HydG, and one GTPase, HydF. The sequence homology between HydG and ThiH, a Radical-SAM enzyme which cleaves tyrosine into p-cresol and dehydroglycine, and the finding of a similar cleavage reaction catalyzed by HydG suggests a mechanism for hydrogenase maturation. Here we propose that HydG is specifically involved in the synthesis of the dithiolate ligand, with two tyrosine-derived dehydroglycines as precursors along with an [FeS] cluster of HydG functioning both as electron shuttle and source of the sulfur atoms.
FEBS Lett
2009;583(3):506-511
| PubMed ID:
19166853
Kuchenreuther JM, Myers WK, Stich TA, George SJ, Nejatyjahromy Y, Swartz JR, Britt RD
A radical intermediate in tyrosine scission to the CO and CN- ligands of FeFe hydrogenase
▸ Abstract
The radical S-adenosylmethionine (SAM) enzyme HydG lyses free l-tyrosine to produce CO and CN(-) for the assembly of the catalytic H cluster of FeFe hydrogenase. We used electron paramagnetic resonance spectroscopy to detect and characterize HydG reaction intermediates generated with a set of (2)H, (13)C, and (15)N nuclear spin-labeled tyrosine substrates. We propose a detailed reaction mechanism in which the radical SAM reaction, initiated at an N-terminal 4Fe-4S cluster, generates a tyrosine radical bound to a C-terminal 4Fe-4S cluster. Heterolytic cleavage of this tyrosine radical at the Cα-Cβ bond forms a transient 4-oxidobenzyl (4OB(•)) radical and a dehydroglycine bound to the C-terminal 4Fe-4S cluster. Electron and proton transfer to this 4OB(•) radical forms p-cresol, with the conversion of this dehydroglycine ligand to Fe-bound CO and CN(-), a key intermediate in the assembly of the 2Fe subunit of the H cluster.
Science
2013;342(6157):472-475
| PubMed ID:
24159045
Kuchenreuther JM, Myers WK, Suess DL, Stich TA, Pelmenschikov V, Shiigi SA, Cramer SP, Swartz JR, Britt RD, George SJ
The HydG enzyme generates an Fe(CO)2(CN) synthon in assembly of the FeFe hydrogenase H-cluster
▸ Abstract
Three iron-sulfur proteins--HydE, HydF, and HydG--play a key role in the synthesis of the [2Fe](H) component of the catalytic H-cluster of FeFe hydrogenase. The radical S-adenosyl-L-methionine enzyme HydG lyses free tyrosine to produce p-cresol and the CO and CN(-) ligands of the [2Fe](H) cluster. Here, we applied stopped-flow Fourier transform infrared and electron-nuclear double resonance spectroscopies to probe the formation of HydG-bound Fe-containing species bearing CO and CN(-) ligands with spectroscopic signatures that evolve on the 1- to 1000-second time scale. Through study of the (13)C, (15)N, and (57)Fe isotopologs of these intermediates and products, we identify the final HydG-bound species as an organometallic Fe(CO)2(CN) synthon that is ultimately transferred to apohydrogenase to form the [2Fe](H) component of the H-cluster.
Science
2014;343(6169):424-427
| PubMed ID:
24458644
Dinis P, Suess DL, Fox SJ, Harmer JE, Driesener RC, De La Paz L, Swartz JR, Essex JW, Britt RD, Roach PL
X-ray crystallographic and EPR spectroscopic analysis of HydG, a maturase in [FeFe]-hydrogenase H-cluster assembly
▸ Abstract
Hydrogenases use complex metal cofactors to catalyze the reversible formation of hydrogen. In [FeFe]-hydrogenases, the H-cluster cofactor includes a diiron subcluster containing azadithiolate, three CO, and two CN(-) ligands. During the assembly of the H cluster, the radical S-adenosyl methionine (SAM) enzyme HydG lyses the substrate tyrosine to yield the diatomic ligands. These diatomic products form an enzyme-bound Fe(CO)x(CN)y synthon that serves as a precursor for eventual H-cluster assembly. To further elucidate the mechanism of this complex reaction, we report the crystal structure and EPR analysis of HydG. At one end of the HydG (βα)8 triosephosphate isomerase (TIM) barrel, a canonical [4Fe-4S] cluster binds SAM in close proximity to the proposed tyrosine binding site. At the opposite end of the active-site cavity, the structure reveals the auxiliary Fe-S cluster in two states: one monomer contains a [4Fe-5S] cluster, and the other monomer contains a [5Fe-5S] cluster consisting of a [4Fe-4S] cubane bridged by a μ2-sulfide ion to a mononuclear Fe(2+) center. This fifth iron is held in place by a single highly conserved protein-derived ligand: histidine 265. EPR analysis confirms the presence of the [5Fe-5S] cluster, which on incubation with cyanide, undergoes loss of the labile iron to yield a [4Fe-4S] cluster. We hypothesize that the labile iron of the [5Fe-5S] cluster is the site of Fe(CO)x(CN)y synthon formation and that the limited bonding between this iron and HydG may facilitate transfer of the intact synthon to its cognate acceptor for subsequent H-cluster assembly.
Proc Natl Acad Sci U S A
2015;112(5):1362-1367
| PubMed ID:
25605932
Nicolet Y, Pagnier A, Zeppieri L, Martin L, Amara P, Fontecilla-Camps JC
Crystal structure of HydG from Carboxydothermus hydrogenoformans: a trifunctional [FeFe]-hydrogenase maturase
▸ Abstract
The structure of the radical S-adenosyl-L-methionine (SAM) [FeFe]-hydrogenase maturase HydG involved in CN(-) /CO synthesis is characterized by two internal tunnels connecting its tyrosine-binding pocket with the external medium and the C-terminal Fe4 S4 cluster-containing region. A comparison with a tryptophan-bound NosL structure suggests that substrate binding causes the closing of the first tunnel and, along with mutagenesis studies, that tyrosine binds to HydG with its amino group well positioned for H-abstraction by SAM. In this orientation the dehydroglycine (DHG) fragment caused by tyrosine Cα-Cβ bond scission can readily migrate through the second tunnel towards the C-terminal domain where both CN(-) and CO are synthesized. Our HydG structure appears to be in a relaxed state with its C-terminal cluster CysX2 CysX22 Cys motif exposed to solvent. A rotation of this domain coupled to Fe4 S4 cluster assembly would bury its putatively reactive unique Fe ion thereby allowing it to interact with DHG.
Chembiochem
2015;16(3):397-402
| PubMed ID:
25504963
Suess DL, Britt RD
EPR Spectroscopic Studies of [FeFe]-Hydrogenase Maturation
▸ Abstract
Proton reduction and H2 oxidation are key elementary reactions for solar fuel production. Hydrogenases interconvert H(+) and H2 with remarkable efficiency and have therefore received much attention in this context. For [FeFe]-hydrogenases, catalysis occurs at a unique cofactor called the H-cluster. In this article, we discuss ways in which EPR spectroscopy has elucidated aspects of the bioassembly of the H-cluster, with a focus on four case studies: EPR spectroscopic identification of a radical en route to the CO and CN(-) ligands of the H-cluster, tracing (57)Fe from the maturase HydG into the H-cluster, characterization of the auxiliary Fe-S cluster in HydG, and isotopic labeling of the CN(-) ligands of HydA for electronic structure studies of its Hox state. Advances in cell-free maturation protocols have enabled several of these mechanistic studies, and understanding H-cluster maturation may in turn provide insights leading to improvements in hydrogenase production for biotechnological applications.
Catal Letters
2015;58(12):699-707
| PubMed ID:
26508821
Suess DL, Kuchenreuther JM, De La Paz L, Swartz JR, Britt RD
Biosynthesis of the [FeFe] Hydrogenase H Cluster: A Central Role for the Radical SAM Enzyme HydG
▸ Abstract
Hydrogenase enzymes catalyze the rapid and reversible interconversion of H2 with protons and electrons. The active site of the [FeFe] hydrogenase is the H cluster, which consists of a [4Fe-4S]H subcluster linked to an organometallic [2Fe]H subcluster. Understanding the biosynthesis and catalytic mechanism of this structurally unusual active site will aid in the development of synthetic and biological hydrogenase catalysts for applications in solar fuel generation. The [2Fe]H subcluster is synthesized and inserted by three maturase enzymes-HydE, HydF, and HydG-in a complex process that involves inorganic, organometallic, and organic radical chemistry. HydG is a member of the radical S-adenosyl-l-methionine (SAM) family of enzymes and is thought to play a prominent role in [2Fe]H subcluster biosynthesis by converting inorganic Fe(2+), l-cysteine (Cys), and l-tyrosine (Tyr) into an organometallic [(Cys)Fe(CO)2(CN)](-) intermediate that is eventually incorporated into the [2Fe]H subcluster. In this Forum Article, the mechanism of [2Fe]H subcluster biosynthesis is discussed with a focus on how this key [(Cys)Fe(CO)2(CN)](-) species is formed. Particular attention is given to the initial metallocluster composition of HydG, the modes of substrate binding (Fe(2+), Cys, Tyr, and SAM), the mechanism of SAM-mediated Tyr cleavage to CO and CN(-), and the identification of the final organometallic products of the reaction.
Inorg Chem
2015;None(None):None-None
| PubMed ID:
26703931
Pagnier A, Martin L, Zeppieri L, Nicolet Y, Fontecilla-Camps JC
CO and CN- syntheses by [FeFe]-hydrogenase maturase HydG are catalytically differentiated events
▸ Abstract
The synthesis and assembly of the active site [FeFe] unit of [FeFe]-hydrogenases require at least three maturases. The radical S-adenosyl-l-methionine HydG, the best characterized of these proteins, is responsible for the synthesis of the hydrogenase CO and CN(-) ligands from tyrosine-derived dehydroglycine (DHG). We speculated that CN(-) and the CO precursor (-):CO2H may be generated through an elimination reaction. We tested this hypothesis with both wild type and HydG variants defective in second iron-sulfur cluster coordination by measuring the in vitro production of CO, CN(-), and (-):CO2H-derived formate. We indeed observed formate production under these conditions. We conclude that HydG is a multifunctional enzyme that produces DHG, CN(-), and CO at three well-differentiated catalytic sites. We also speculate that homocysteine, cysteine, or a related ligand could be involved in Fe(CO)x(CN)y transfer to the HydF carrier/scaffold.
Proc Natl Acad Sci U S A
2015;None(None):None-None
| PubMed ID:
26699472
Suess DL, Pham CC, Bürstel I, Swartz JR, Cramer SP, Britt RD
The Radical SAM Enzyme HydG Requires Cysteine and a Dangler Iron for Generating an Organometallic Precursor to the [FeFe]-Hydrogenase H-Cluster
▸ Abstract
Three maturase enzymes-HydE, HydF, and HydG-synthesize and insert the organometallic component of the [FeFe]-hydrogenase active site (the H-cluster). HydG generates the first organometallic intermediates in this process, ultimately producing an [Fe(CO)2(CN)] complex. A limitation in understanding the mechanism by which this complex forms has been uncertainty regarding the precise metallocluster composition of HydG that comprises active enzyme. We herein show that the HydG auxiliary cluster must bind both l-cysteine and a dangler Fe in order to generate the [Fe(CO)2(CN)] product. These findings support a mechanistic framework in which a [(Cys)Fe(CO)2(CN)](-) species is a key intermediate in H-cluster maturation.
J Am Chem Soc
2016;138(4):1146-1149
| PubMed ID:
26764535