Top Level Name
⌊ Superfamily (core) Haloacid Dehalogenase
⌊ Subgroup C1.8: polynucleotide 5'-hydroxyl-kinase c-terminal phosphatase like
⌊ Family polynucleotide 5'-hydroxyl-kinase c-terminal phosphatase
| Total |
100%  |
<100%  |
|||
| Functional domains | 1 | 0 | 1 | ||
| UniProtKB | 1 | 0 | 1 | ||
| GI | 4 | 0 | 4 | ||
| Structures | 5 | ||||
| Reactions | 1 | ||||
| Functional domains of this family were last updated on Nov. 22, 2017 | |||||
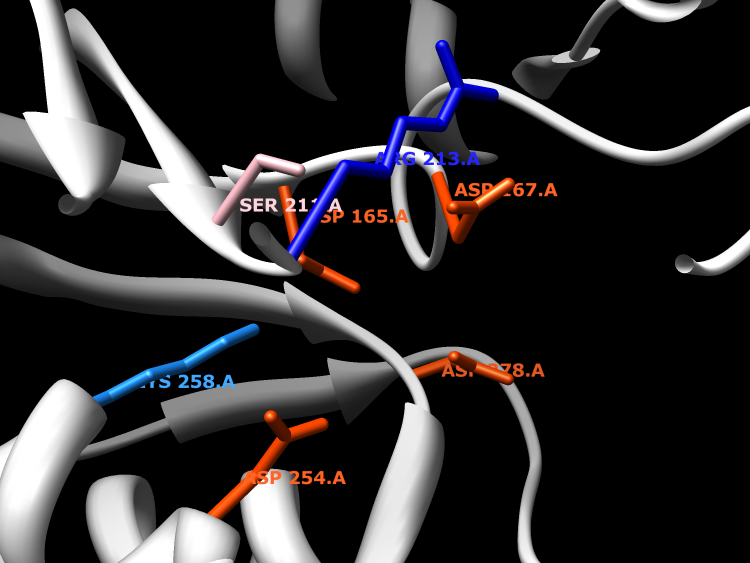
Enzymes in the polynucleotide 5'-hydroxyl-kinase c-terminal phosphatase family contain two distinct structural domains, each with a seperate active site. Only the C-terminal domain is part of the haloacid dehalogenase superfamily. This domain has 3'-phosphatase activity, and can hydrolyze 2'3'-cyclic phosphodiesters. However, the biologically relevant phosphatase reaction has not yet been identified. Mg2+ is required as a cofactor. The N-terminal domain has kinase activity, but is not a part of the haloacid dehalogenase superfamily.
Galburt, E.A., et al.
Structure of a tRNA repair enzyme and molecular biology workhorse: T4 polynucleotide kinase
▸ Abstract
Structure (Camb) 2002;10(9):1249-1260 | PubMed ID: 12220496
No notes.
Active Site
Catalyzed Reaction(s)
polynucleotide 5'-hydroxyl-kinase
 |
+ | 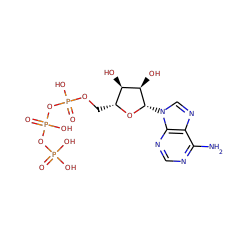 |
 |
 |
+ | 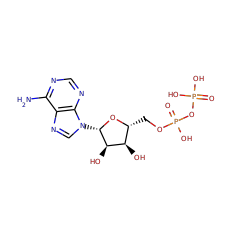 |
||
| 5'-dephospho-DNA 16678 |
ATP 15422 |
5'-phospho-DNA 16149 |
ADP 16761 |
EC: | IntEnz: | Kegg: | BioCyc: | BRENDA: |