Top Level Name
⌊ Superfamily (core) Haloacid Dehalogenase
⌊ Subgroup C1.5: HAD, Beta-PGM, Phosphatase Like
⌊ C1.5.6: HAD, Beta-PGM, Phosphatase Like
⌊ FunctionalDomain C1.5.6: HAD, Beta-PGM, Phosphatase Like (ID 116721)
No Notes.
| Superfamily Assignment Evidence Code(s) | ISS |
| This entry was last updated on | Nov. 22, 2017 |
References to Other Databases
Genbank
| Species | GI | Accession | Proteome |
|---|---|---|---|
| Rattus norvegicus Taxon ID: 10116 | 57863283 | NP_001009409.1 (RefSeq) | PRP URP |
| Rattus norvegicus Taxon ID: 10116 | 149031100 | EDL86127.1 (Genbank) | PRP URP |
| Rattus norvegicus Taxon ID: 10116 | 56541034 | AAH87587.1 (Genbank) | PRP URP |
| Rattus norvegicus Taxon ID: 10116 | 81883158 | PRP URP | |
| Show All | |||
Uniprot
| Protein Name | Accession | EC Number
 |
Identifier |
|---|---|---|---|
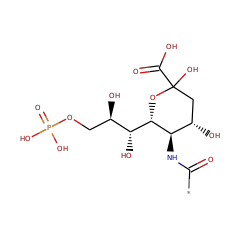
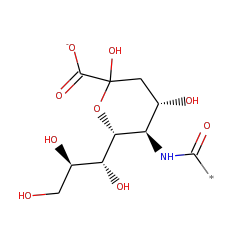
| N-acylneuraminate-9-phosphatase | Q5M969 | 3.1.3.29 | NANP_RAT (Swiss-Prot) |
Length of Enzyme (full-length): 248 | Length of Functional Domain: 238
MGLSRVRAVFFDLDNTLIDTAGASRRGMLEVIKLLQSKYHYKEEAEVICDKVQVKLSKEC
FHPYSTCITDVRTSHWEEAIQETKGGADNRKLAEECYFLWKSTRLQHMTLEEDVKAMLTE
LRKEVRLLLLTNGDRQTQREKIEACACQSYFDAIVVGGEQKEEKPAPSIFYHCCDLLGVQ
PGDCVMVGDTLETDIQGGLNAGLKATVWINKSGGVPLTSSPMPHYMVSSVLELPALLQSI
DCKVSMSV
Conserved catalytic residues (as determined by automated alignment to family, subgroup, or superfamily HMMs) are shown with teal highlighting . Conserved catalytic residues which do not matched the Conserved Alignment Residue are shown with maroon highlighting . Information regarding their function can be found in the Conserved Residues section below.