Family known  |
|||||||
| Total |
100%  |
<100%  |
Family unknown  |
||||
| Functional domains | 25354 | 0 | 19755 | 5599 | |||
| UniProtKB | 86575 | 0 | 77128 | 9447 | |||
| GI | 159419 | 0 | 140832 | 18587 | |||
| Structures | 135 | ||||||
| Reactions | 6 | ||||||
| Functional domains of this subgroup were last updated on Nov. 22, 2017 | |||||||
| New functional domains were last added to this subgroup on Aug. 1, 2014 | |||||||
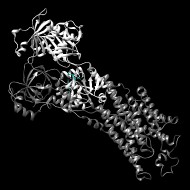
Experimentally characterized enzymes in the P-type ATPase subgroup harness the energy of ATP to pump charged substrates across biological membranes. Mg2+ is required as a cofactor. ATPases in this group transport various molecules including heavy metals, phospholipids, K+, Ca2+, H+, Mg2+, Na+/K+, and H+/K+. P-type ATPases are made up of multiple structural domains. Only the catalytic domain, responsible for the hydrolysis of ATP, is part of the haloacid dehalogenase (HAD) superfamily.
Toyoshima, C., et al.
Crystal structure of the calcium pump of sarcoplasmic reticulum at 2.6 A resolution
▸ Abstract
Nature 2000;405(6787):647-655 | PubMed ID: 10864315
Axelsen, K.B. and M.G. Palmgren
Evolution of substrate specificities in the P-type ATPase superfamily
▸ Abstract
J Mol Evol 1998;46(1):84-101 | PubMed ID: 9419228
Aravind L, Galperin MY, Koonin EV
The catalytic domain of the P-type ATPase has the haloacid dehalogenase fold
▸ Abstract
Trends Biochem Sci 1998;23(4):127-129 | PubMed ID: 9584613
No notes.
Static File Downloads
| File Name | Description | Parameters | Stats |
|---|---|---|---|
| repnet.sg2.th50.pE20.mek250.xgmml | Representative network: each node is a group of similar sequences | node similarity threshold = 50 max edge count = 250 min -log10 E = 20 |
size = 86M num_edges = 250000 num_nodes = 2797 |
| sfld_alignment_sg2.msa | Annotated Sequence Alignment, Stockholm format | 38 sequences size: 53K |
| Subgroup ▸ Legend | T  |
K  |
C  |
U  |
S  |
||
|---|---|---|---|---|---|---|---|
| C1.7: P-type atpase like | 25354 | 19755 | 0 | 5599 | 135 | ||
| ┗ p-type atpase | 19755 | 19755 | 0 | 99 |