Family known  |
|||||||
| Total |
100%  |
<100%  |
Family unknown  |
||||
| Functional domains | 139 | 0 | 0 | 139 | |||
| UniProtKB | 128 | 0 | 0 | 128 | |||
| GI | 367 | 0 | 0 | 367 | |||
| Structures | 15 | ||||||
| Reactions | 0 | ||||||
| Functional domains of this subgroup were last updated on Nov. 22, 2017 | |||||||
| New functional domains were last added to this subgroup on Aug. 1, 2014 | |||||||
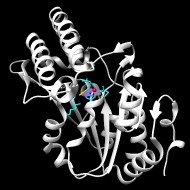
Structurally characterized enzymes in this subgroup have the C1 cap type
Bitto E, Bingman CA, Wesenberg GE, McCoy JG, Phillips GN
Structure of pyrimidine 5'-nucleotidase type 1. Insight into mechanism of action and inhibition during lead poisoning
▸ Abstract
J Biol Chem 2006;281(29):20521-20529 | PubMed ID: 16672222
No notes.
Static File Downloads
| File Name | Description | Parameters | Stats |
|---|---|---|---|
| network.sg1128.bs60.mek250K.xgmml | One node per sequence network | min bit score = 60 max edge count = 250K |
size = 4.5M num_edges = 8603 num_nodes = 139 |
| repnet.sg1128.th50.pE20.mek250.xgmml | Representative network: each node is a group of similar sequences | node similarity threshold = 50 max edge count = 250 min -log10 E = 20 |
size = 183K num_edges = 303 num_nodes = 34 |
| sfld_alignment_sg1128.msa | Annotated Sequence Alignment, Stockholm format | 60 sequences size: 33K |
Total number of functional domains in this group.
Number of Functional Domains that have been manually or automatically been assigned to a family.
Number of Functional Domains that have not been assigned to a family.
Number of structures available from the PDB for members of this group.
Number of Functional Domains with 100% of Conserved Residues
Number of Functional Domains with less than 100% Conserved Residues
| Subgroup ▸ Legend | T  |
K  |
C  |
U  |
S  |
||
|---|---|---|---|---|---|---|---|
| C1.4: 5'-Nucleotidase Like | 139 | 0 | 0 | 139 | 15 |
Depth of the multi-level Subgroup hierarchy.