Ruszczycky MW, Ogasawara Y, Liu HW
Radical SAM enzymes in the biosynthesis of sugar-containing natural products
▸ Abstract
Carbohydrates play a key role in the biological activity of numerous natural products. In many instances their biosynthesis requires radical mediated rearrangements, some of which are catalyzed by radical SAM enzymes. BtrN is one such enzyme responsible for the dehydrogenation of a secondary alcohol in the biosynthesis of 2-deoxystreptamine. DesII is another example that catalyzes a deamination reaction necessary for the net C4 deoxygenation of a glucose derivative en route to desosamine formation. BtrN and DesII represent the two most extensively characterized radical SAM enzymes involved in carbohydrate biosynthesis. In this review, we summarize the biosynthetic roles of these two enzymes, their mechanisms of catalysis, the questions that have arisen during these investigations and the insight they can offer for furthering our understanding of radical SAM enzymology. This article is part of a Special Issue entitled: Radical SAM enzymes and Radical Enzymology.
Biochim Biophys Acta
2012;1824(11):1231-1244
| PubMed ID:
22172915
Yokoyama K, Numakura M, Kudo F, Ohmori D, Eguchi T
Characterization and mechanistic study of a radical SAM dehydrogenase in the biosynthesis of butirosin
▸ Abstract
BtrN encoded in the butirosin biosynthetic gene cluster possesses a CXXXCXXC motif conserved within the radical S-adenosyl methionine (SAM) superfamily. Its gene disruption in the butirosin producer Bacillus circulans caused the interruption of the biosynthetic pathway between 2-deoxy-scyllo-inosamine (DOIA) and 2-deoxystreptamine (DOS). Further, in vitro assay of the overexpressed enzyme revealed that BtrN catalyzed the oxidation of DOIA under the strictly anaerobic conditions along with consumption of an equimolar amount of SAM to produce 5'-deoxyadenosine, methionine, and 3-amino-2,3-dideoxy-scyllo-inosose (amino-DOI). Kinetic analysis showed substrate inhibition by DOIA but not by SAM, which suggests that the reaction is the Ordered Bi Ter mechanism and that SAM is the first substrate and DOIA is the second. The BtrN reaction with [3-2H]DOIA generated nonlabeled, monodeuterated and dideuterated 5'-deoxyadenosines, while no deuterium was incorporated by incubation of nonlabeled DOIA in the deuterium oxide buffer. These results indicated that the hydrogen atom at C-3 of DOIA was directly transferred to 5'-deoxyadenosine to give the radical intermediate of DOIA. Generation of nonlabeled and dideuterated 5'-deoxyadenosines proved the reversibility of the hydrogen abstraction step. The present study suggests that BtrN is an unusual radical SAM dehydrogenase catalyzing the oxidation of the hydroxyl group by a radical mechanism. This is the first report of the mechanistic study on the oxidation of a hydroxyl group by a radical SAM enzyme.
J Am Chem Soc
2007;129(49):15147-15155
| PubMed ID:
18001019
Goldman PJ, Grove TL, Booker SJ, Drennan CL
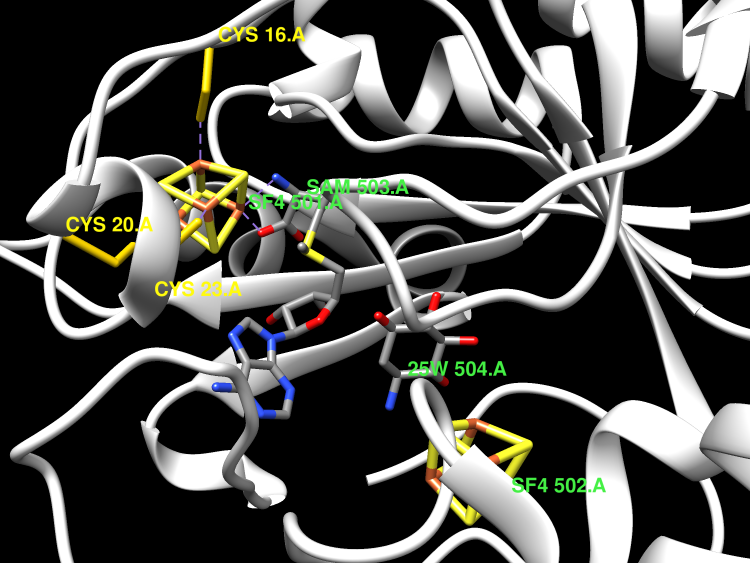
X-ray analysis of butirosin biosynthetic enzyme BtrN redefines structural motifs for AdoMet radical chemistry
▸ Abstract
The 2-deoxy-scyllo-inosamine (DOIA) dehydrogenases are key enzymes in the biosynthesis of 2-deoxystreptamine-containing aminoglycoside antibiotics. In contrast to most DOIA dehydrogenases, which are NAD-dependent, the DOIA dehydrogenase from Bacillus circulans (BtrN) is an S-adenosyl-l-methionine (AdoMet) radical enzyme. To examine how BtrN employs AdoMet radical chemistry, we have determined its structure with AdoMet and substrate to 1.56 Å resolution. We find a previously undescribed modification to the core AdoMet radical fold: instead of the canonical (β/α)6 architecture, BtrN displays a (β5/α4) motif. We further find that an auxiliary [4Fe-4S] cluster in BtrN, thought to bind substrate, is instead implicated in substrate-radical oxidation. High structural homology in the auxiliary cluster binding region between BtrN, fellow AdoMet radical dehydrogenase anSME, and molybdenum cofactor biosynthetic enzyme MoaA provides support for the establishment of an AdoMet radical structural motif that is likely common to ∼6,400 uncharacterized AdoMet radical enzymes.
Proc Natl Acad Sci U S A
2013;110(40):15949-15954
| PubMed ID:
24048029
Maiocco SJ, Grove TL, Booker SJ, Elliott SJ
Electrochemical Resolution of the [4Fe-4S] Centers of the AdoMet Radical Enzyme BtrN: Evidence of Proton Coupling and an Unusual, Low-Potential Auxiliary Cluster
▸ Abstract
The S-adenosylmethionine (AdoMet) radical superfamily of enzymes includes over 113 500 unique members, each of which contains one indispensable iron-sulfur (FeS) cluster that is required to generate a 5'-deoxyadenosyl 5'-radical intermediate during catalysis. Enzymes within several subgroups of the superfamily, however, have been found to contain one or more additional FeS clusters. While these additional clusters are absolutely essential for enzyme activity, their exact roles in the function and/or mechanism of action of many of the enzymes are at best speculative, indicating a need to develop methods to characterize and study these clusters in more detail. Here, BtrN, an AdoMet radical dehydrogenase that catalyzes the two-electron oxidation of 2-deoxy-scyllo-inosamine to amino-dideoxy-scyllo-inosose, an intermediate in the biosynthesis of 2-deoxystreptamine antibiotics, is examined through direct electrochemistry, where the potential of both its AdoMet radical and auxiliary [4Fe-4S] clusters can be measured simultaneously. We find that the AdoMet radical cluster exhibits a midpoint potential of -510 mV, while the auxiliary cluster exhibits a midpoint potential of -765 mV, to our knowledge the lowest [4Fe-4S](2+/+) potential to be determined to date. The impact of AdoMet binding and the pH dependence of catalysis are also quantitatively observed. These data show that direct electrochemical methods can be used to further elucidate the chemistry of the burgeoning AdoMet radical superfamily in the future.
J Am Chem Soc
2015;137(27):8664-8667
| PubMed ID:
26088836
Grove, Ahlum, Sharma, Krebs & Booker
A consensus mechanism for Radical SAM-dependent dehydrogenation? BtrN contains two [4Fe-4S] clusters.
▸ Abstract
BtrN catalyzes the two-electron oxidation of the C3 secondary alcohol of 2-deoxy-scyllo-inosamine to the corresponding ketone and is a member of a subclass of radical S-adenosylmethionine (SAM) enzymes called radical SAM (RS) dehydrogenases. Like all RS enzymes, BtrN contains a [4Fe-4S] cluster that delivers an electron to SAM, inducing its cleavage to the common intermediate in RS reactions, the 5'-deoxyadenosyl 5'-radical. In this work, we show that BtrN contains an additional [4Fe-4S] cluster, thought to bind in contact with the substrate to facilitate loss of the second electron in the two-electron oxidation.
Biochemistry
2010;49(18):3783-3785
| PubMed ID:
20377206
Yokoyama, Ohmori, Kudo & Eguchi
Mechanistic study on the reaction of a radical SAM dehydrogenase BtrN by electron paramagnetic resonance spectroscopy
▸ Abstract
BtrN is a radical SAM ( S-adenosyl- l-methionine) enzyme that catalyzes the oxidation of 2-deoxy- scyllo-inosamine (DOIA) into 3-amino-2,3-dideoxy- scyllo-inosose (amino-DOI) during the biosynthesis of 2-deoxystreptamine (DOS) in the butirosin producer Bacillus circulans. Recently, we have shown that BtrN catalyzes the transfer of a hydrogen atom at C-3 of DOIA to 5'-deoxyadenosine, and thus, the reaction was proposed to proceed through the hydrogen atom abstraction by the 5'-deoxyadenosyl radical. In this work, the BtrN reaction was analyzed by EPR spectroscopy. A sharp double triplet EPR signal was observed when the EPR spectrum of the enzyme reaction mixture was recorded at 50 K. The spin coupling with protons partially disappeared by reaction with [2,2- (2)H 2]DOIA, which unambiguously proved the observed signal to be a radical on C-3 of DOIA. On the other hand, the EPR spectrum of the [4Fe-4S] cluster of BtrN during the reaction showed a complex signal due to the presence of several species. Comparison of signals derived from a [4Fe-4S] center of BtrN incubated with various combinations of products (5'-deoxyadenosine, l-methionine, and amino-DOI) and substrates (SAM and DOIA) indicated that the EPR signals observed during the reaction were derived from free BtrN, a BtrN-SAM complex, and a BtrN-SAM-DOIA complex. Significant changes in the EPR signals upon binding of SAM and DOIA suggest the close interaction of both substrates with the [4Fe-4S] cluster.
Bichemistry
2008;47(34):8950-8960
| PubMed ID:
18672902