Top Level Name
⌊ Superfamily (core) Enolase
⌊ Subgroup mandelate racemase
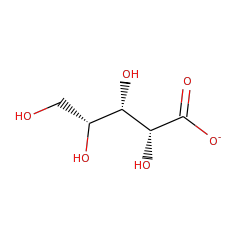
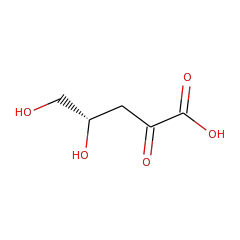
⌊ Family xylonate dehydratase 1
⌊ FunctionalDomain xylonate dehydratase 1 (ID 50)
No Notes.
| Superfamily Assignment Evidence Code(s) | ISS |
| Family Assignment Evidence Code | IES PubMed:20736170 |
| This entry was last updated on | Nov. 21, 2017 |
References to Other Databases
Genbank
| Species | GI | Accession | Proteome |
|---|---|---|---|
| Sulfolobus solfataricus Taxon ID: 2287 | 497674398 | WP_009988582.1 (RefSeq) | URP |
| Sulfolobus solfataricus Taxon ID: 2287 | 800906085 | AKA78266.1 (Genbank) | URP |
| Sulfolobus solfataricus Taxon ID: 2287 | 800903391 | AKA75573.1 (Genbank) | URP |
| Sulfolobus solfataricus Taxon ID: 2287 | 800900691 | AKA72874.1 (Genbank) | URP |
| Sulfolobus solfataricus 98/2 Taxon ID: 555311 | 261601155 | ACX90758.1 (Genbank) | URP |
| Sulfolobus solfataricus P2 Taxon ID: 273057 | 13815980 | AAK42783.1 (Genbank) | PRP URP |
| obsolete GIs = 284173189, 384433001, 15899388 | |||
| Show All | |||
Uniprot
| Protein Name | Accession | EC Number
 |
Identifier |
|---|---|---|---|
| n/a | A0A0E3JWC5 | A0A0E3JWC5_SULSF (TrEMBL) | |
| n/a | D0KNY8 | D0KNY8_SULS9 (TrEMBL) | |
| n/a | Q97VG1 | Q97VG1_SULSO (TrEMBL) |
Length of Enzyme (full-length): 393 | Length of Functional Domain: 393
MTKISEIEAYILGKEVTSAQWASLMVLVRVTTNDGRVGWGETVSALRAEAVANFVKKINT
VLKGNDVFNVEKNRLEWYKHDFNMTISLESTTAYSAVDIASWDIIGKELGAPLYKLLGGK
TRDKVLVYANGWYQNCVKPEDFAEKAKEIVKMGYKALKFDPFGPYFNDISKKGLDIAEER
VKAVREAVGDNVDILIEHHGRFNANSAIMIAKRLEKYNPLFMEEPIHPEDVEGLRKYRNN
TSLRIALGERIINKQQALYFMKEGLVDFLQADLYRIGGVTETKKVVGIAETFDVQMAFHN
AQGPILNAVTLQFDAFIPNFLIQESFYDWFPSWKRELIYNGTPIDNGYAIIPERPGLGVE
VNEKMLDSLKVKGEEYFNPEEPVWVVKGTWRDY
Conserved catalytic residues (as determined by automated alignment to family, subgroup, or superfamily HMMs) are shown with teal highlighting . Conserved catalytic residues which do not matched the Conserved Alignment Residue are shown with maroon highlighting . Information regarding their function can be found in the Conserved Residues section below.