Top Level Name
⌊ Superfamily (core) Radical SAM
⌊ Subgroup SPASM/twitch domain containing
⌊ Family UDP-N-acetyl-tunicamine-uracil synthase (TunB-like)
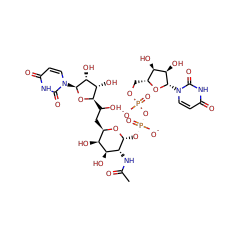
⌊ FunctionalDomain UDP-N-acetyl-tunicamine-uracil synthase (TunB) (ID 434568)
No Notes.
| Superfamily Assignment Evidence Code(s) | ISS |
| Family Assignment Evidence Code | CFM PubMed:22172915 |
| This entry was last updated on | June 10, 2017 |
References to Other Databases
Genbank
| Species | GI | Accession | Proteome |
|---|---|---|---|
| Streptomyces chartreusis Taxon ID: 1969 | 497731628 | WP_010045812.1 (RefSeq) | |
| Streptomyces chartreusis NRRL 3882 Taxon ID: 1079985 | 341904718 | AEL00521.1 (Genbank) | |
| Streptomyces chartreusis NRRL 3882 Taxon ID: 1079985 | 334903193 | AEH25658.1 (Genbank) | |
| Streptomyces chartreusis NRRL 3882 Taxon ID: 1079985 | 311404562 | ADP94222.1 (Genbank) | |
| obsolete GI = 383650027 | |||
| Show All | |||
Uniprot
| Protein Name | Accession | EC Number
 |
Identifier |
|---|---|---|---|
| n/a | E5KJ95 | E5KJ95_STRCX (TrEMBL) |
Length of Enzyme (full-length): 338 | Length of Functional Domain: 338
MTGYTAPVRASLDVTYACDLRCIHCRTNTGEIPVHIRRKMLSIEQLQDIMLQLDRMGTFE
ITLTGGEPTIRKGFWQLLDVVPKLRHSTVTLITNAAAHTREQLDRIVDSGISSIRVSMDG
TRETFAAVRLMDVFDTVVENSIYLRDRVRSFKVLTTVMKTNQHNVFDLAHYLRAHGFRRQ
DLILVRAHGRGGRNRLLLTEEETMAVHRKVTEFKRDVPATDFDLNLNAPYLVPGEKLRPF
QDVVMFPYLVRDSSIAISATGDVTMSRLYSAAPLGNTKDAPVQEIWTQGQQQLVQEQEEF
TEERLREIFWDFATSDGPDGPPLTSLLDRQIFEGAEVR
Conserved catalytic residues (as determined by automated alignment to family, subgroup, or superfamily HMMs) are shown with teal highlighting . Conserved catalytic residues which do not matched the Conserved Alignment Residue are shown with maroon highlighting . Information regarding their function can be found in the Conserved Residues section below.