Benjdia A, Subramanian S, Leprince J, Vaudry H, Johnson MK, Berteau O
Anaerobic sulfatase-maturating enzymes, first dual substrate radical S-adenosylmethionine enzymes
▸ Abstract
Sulfatases are a major group of enzymes involved in many critical physiological processes as reflected by their broad distribution in all three domains of life. This class of hydrolases is unique in requiring an essential post-translational modification of a critical active-site cysteine or serine residue to C(alpha)-formylglycine. This modification is catalyzed by at least three nonhomologous enzymatic systems in bacteria. Each enzymatic system is currently considered to be dedicated to the modification of either cysteine or serine residues encoded in the sulfatase-active site and has been accordingly categorized as Cys-type and Ser-type sulfatase-maturating enzymes. We report here the first detailed characterization of two bacterial anaerobic sulfatase-maturating enzymes (anSMEs) that are physiologically responsible for either Cys-type or Ser-type sulfatase maturation. The activity of both enzymes was investigated in vivo and in vitro using synthetic substrates and the successful purification of both enzymes facilitated the first biochemical and spectroscopic characterization of this class of enzyme. We demonstrate that reconstituted anSMEs are radical S-adenosyl-l-methionine enzymes containing a redox active [4Fe-4S](2+,+) cluster that initiates the radical reaction by binding and reductively cleaving S-adenosyl-l-methionine to yield 5 '-deoxyadenosine and methionine. Surprisingly, our results show that anSMEs are dual substrate enzymes able to oxidize both cysteine and serine residues to C(alpha)-formylglycine. Taken together, the results support a radical modification mechanism that is initiated by hydrogen abstraction from a serine or cysteine residue located in an appropriate target sequence.
J Biol Chem.
2008;283(26):17815-17826
| PubMed ID:
18408004
Benjdia A, Leprince J, Guillot A, Vaudry H, Rabot S, Berteau O
Anaerobic sulfatase-maturating enzymes: radical SAM enzymes able to catalyze in vitro sulfatase post-translational modification
▸ Abstract
Sulfatases are widespread enzymes, found from prokaryotes to eukaryotes and involved in many biochemical processes. To be active, all known sulfatases undergo a unique post-translational modification leading to the conversion of a critical active-site residue, i.e., a serine or a cysteine, into a Cα-formylglycine (FGly). Two different systems are involved in sulfatase maturation. One, named FGE, is an oxygen-dependent oxygenase and has been fully characterized. The other one, a member of the so-called “radical SAM” super-family, has been only preliminary investigated. This latter system allows the maturation of sulfatases in strictly anaerobic conditions and has thus been named anSME (anaerobic Sulfatase Maturating Enzyme). Our results provide the first experimental evidence that anSME are iron−sulfur enzymes able to perform the reductive cleavage of SAM and thus belong to the radical SAM super-family. Furthermore, they demonstrate that anSME are able to efficiently oxidize cysteine into FGly in an oxygen-independent manner.
J Am Chem Soc
2007;129(12):3462-3463
| PubMed ID:
17335281
Benjdia A, Subramanian S, Leprince J, Vaudry H, Johnson MK, Berteau O.
Anaerobic sulfatase-maturating enzyme--a mechanistic link with glycyl radical-activating enzymes?
▸ Abstract
Sulfatases form a major group of enzymes present in prokaryotes and eukaryotes. This class of hydrolases is unique in requiring essential post-translational modification of a critical active-site cysteinyl or seryl residue to C(alpha)-formylglycine (FGly). Herein, we report mechanistic investigations of a unique class of radical-S-adenosyl-L-methionine (AdoMet) enzymes, namely anaerobic sulfatase-maturating enzymes (anSMEs), which catalyze the oxidation of Cys-type and Ser-type sulfatases and possess three [4Fe-4S](2+,+) clusters. We were able to develop a reliable quantitative enzymatic assay that allowed the direct measurement of FGly production and AdoMet cleavage. The results demonstrate stoichiometric coupling of AdoMet cleavage and FGly formation using peptide substrates with cysteinyl or seryl active-site residues. Analytical and EPR studies of the reconstituted wild-type enzyme and cysteinyl cluster mutants indicate the presence of three almost isopotential [4Fe-4S](2+,+) clusters, each of which is required for the generation of FGly in vitro. More surprisingly, our data indicate that the two additional [4Fe-4S](2+,+) clusters are required to obtain efficient reductive cleavage of AdoMet, suggesting their involvement in the reduction of the radical AdoMet [4Fe-4S](2+,+) center. These results, in addition to the recent demonstration of direct abstraction by anSMEs of the C(beta) H-atom from the sulfatase active-site cysteinyl or seryl residue using a 5'-deoxyadenosyl radical, provide new insights into the mechanism of this new class of radical-AdoMet enzymes.
FEBS J
1920;2012(277):8-1906
| PubMed ID:
20218986
Goldman PJ, Grove TL, Sites LA, McLaughlin MI, Booker SJ, Drennan CL
X-ray structure of an AdoMet radical activase reveals an anaerobic solution for formylglycine posttranslational modification
▸ Abstract
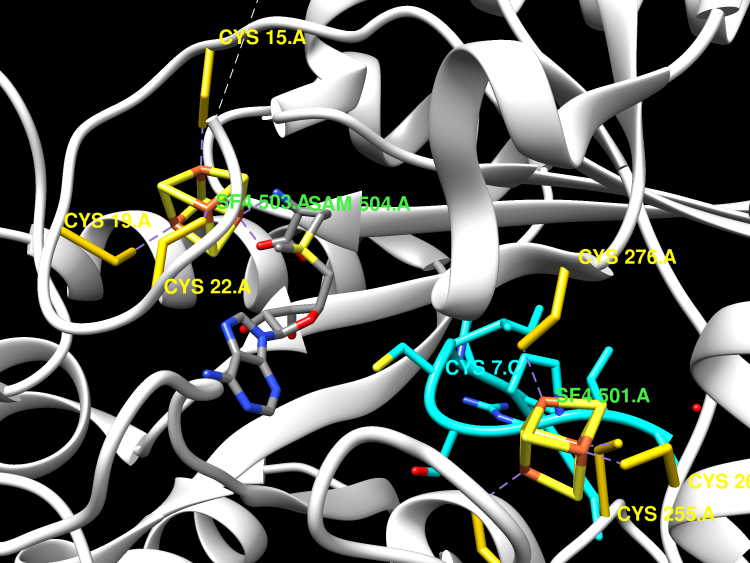
Arylsulfatases require a maturating enzyme to perform a co- or posttranslational modification to form a catalytically essential formylglycine (FGly) residue. In organisms that live aerobically, molecular oxygen is used enzymatically to oxidize cysteine to FGly. Under anaerobic conditions, S-adenosylmethionine (AdoMet) radical chemistry is used. Here we present the structures of an anaerobic sulfatase maturating enzyme (anSME), both with and without peptidyl-substrates, at 1.6-1.8 Å resolution. We find that anSMEs differ from their aerobic counterparts in using backbone-based hydrogen-bonding patterns to interact with their peptidyl-substrates, leading to decreased sequence specificity. These anSME structures from Clostridium perfringens are also the first of an AdoMet radical enzyme that performs dehydrogenase chemistry. Together with accompanying mutagenesis data, a mechanistic proposal is put forth for how AdoMet radical chemistry is coopted to perform a dehydrogenation reaction. In the oxidation of cysteine or serine to FGly by anSME, we identify D277 and an auxiliary [4Fe-4S] cluster as the likely acceptor of the final proton and electron, respectively. D277 and both auxiliary clusters are housed in a cysteine-rich C-terminal domain, termed SPASM domain, that contains homology to ~1,400 other unique AdoMet radical enzymes proposed to use [4Fe-4S] clusters to ligate peptidyl-substrates for subsequent modification. In contrast to this proposal, we find that neither auxiliary cluster in anSME bind substrate, and both are fully ligated by cysteine residues. Instead, our structural data suggest that the placement of these auxiliary clusters creates a conduit for electrons to travel from the buried substrate to the protein surface.
Proc Natl Acad Sci U S A
2013;110(21):8519-8524
| PubMed ID:
23650368
Benjdia A, Berteau O
Sulfatases and radical SAM enzymes: emerging themes in glycosaminoglycan metabolism and the human microbiota
▸ Abstract
Humans live in a permanent association with bacterial populations collectively called the microbiota. In the last 10 years, major advances in our knowledge of the microbiota have shed light on its critical roles in human physiology. The microbiota has also been shown to be a major factor in numerous pathologies including obesity or inflammatory disorders. Despite tremendous progresses, our understanding of the key functions of the human microbiota and the molecular basis of its interactions with the host remain still poorly understood. Among the factors involved in host colonization, two enzymes families, sulfatases and radical S-adenosyl-L-methionine enzymes, have recently emerged as key enzymes.
Biochem Soc Trans
2016;44(1):109-115
| PubMed ID:
26862195